online learning
Psiflow allows for the seamless development and scalable
execution of online learning algorithms for ML potentials.
The Learning class provides an interface based on which such
algorithms can be implemented.
They keep track of the generated data, error metrics, optional Weights &
Biases logging, and provide basic restart functionalities in case
something goes wrong.
Learning objects are instantiated using the following arguments:
- reference (type
Reference): theReferenceinstance which will be used to evaluate ground-truth energy/force labels for each of the samples generated. - path_output (type
str | Path): the location to a folder in which intermediate models, datasets, walker states, and restart files can be saved. - train_valid_split (type
float): fraction of generated data which should be used for the training set (as opposed to validation). - error_thresholds_for_reset (type
list[Optional[float]]): during online learning, it is not uncommon to have walkers explore unphysical regions in phase space due to irregularities in the intermediate potential, excessive temperatures/pressures, ... In those cases, it is beneficial to reset walkers to their starting configurations, of which it is known to be a physically sound starting point. The decision to reset walkers is made every time the 'exact' energy and forces have been computed from a sampled state. If the error between the corresponding walker's model (i.e. the previous model) and the QM-evaluated energy and forces exceeds a certain threshold (both on energies and forces), the walker is reset. This argument expects a list of length two (threshold on energy error, and threshold on force error), with optionalNonevalues if no reset is desired. For example:[None, 0.1]indicates to reset whenever the force RMSE exceeds 100 meV/A, and ignore any energy discrepancy. - error_thresholds_for_discard (type
list[Optional[float]]): states which are entirely unphysical do not contribute to the accuracy of the model, and sometimes even hinder proper training. If these error thresholds are exceeded, the state is discarded and the walker is reset. - wandb_group (type
str): if specified, the computed dataset metrics will be logged to Weights & Biases in the corresponding group of runs for easy visual analysis. - wandb_project (type
str): if specified, the computed dataset metrics will be logged to Weights & Biases in the corresponding project for easy visual analysis. - initial_data (type
Dataset): existing, labeled data from which the learning can be bootstrapped. Note that all states in this dataset must be labeled, and that this is only sensible if the labeling agrees with the given Reference instance. (Same level of theory, same basis set, grid settings, ... ).
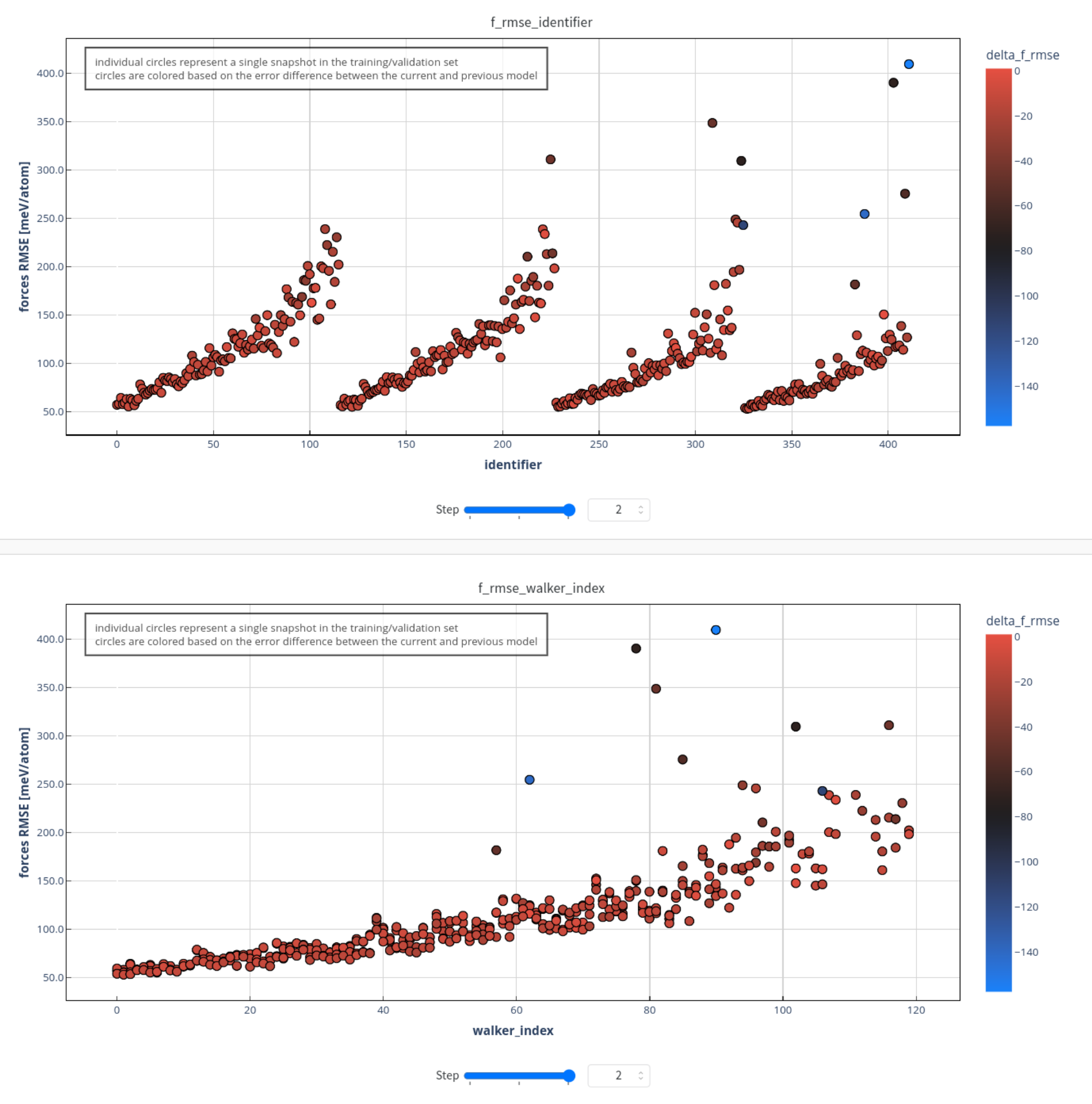
The core business of a Learning instance is the following sequence of operations:
- use walkers in a
sample()call to generate atomic geometries - evaluate those atomic geometries with the provided reference to obtain QM energy and forces
- include those geometries to the training data, or discard them if they exceed
error_thresholds_for_discard. Reset walkers if they exceederror_thresholds_for_reset. - Train the model using the new data.
- Compute metrics for the trained model across the new dataset and optionally log them to W&B.
Currently, there are two variants of this implemented: passive and active learning.
passive learning
During passive learning, walkers are propagated using an external and 'fixed' Hamiltonian which is not trained at any point (e.g. a pre-trained universal potential or a hessian-based Hamiltonian).
model, walkers = learning.passive_learning(
model,
walkers,
hamiltonian=MACEHamiltonian.mace_mp0(), # fixed hamiltonian
steps=20000,
step=2000,
**optional_sampling_kwargs,
)
passive_learning()
call.
Additional keyword arguments to this function are passed directly into the sample function (e.g. for
specifying the log level or the center-of-mass behavior).
The returned model is the one trained on all data generated in the passive_learning() call as well as all data which was already present in the learning instance (for example if it had been initialized with initial_data, see above).
The returned walkers are identical to the ones passed into the method, but this is done to
emphasize that internally, they do change due to calling passive_learning (because they
are either propagated or reset, or their metadynamics bias has changed because there are
more hills present than before).
active learning
During active learning, walkers are propagated with a Hamiltonian generated using the current model. They are propagated for a given number of steps after which their final state is passed into the reference for correct labeling. Different from passive learning, active learning does not allow for subsampling of the trajectories of the walkers. The idea behind this is that if you wish to propagate the walker for 10 ps, and sample a structure every 1 ps to let each walker generate 10 states, it is likely much better to instead increase the number of walkers (to cover more regions in phase space) and propagate them in steps of 1 ps. Active learning is ideally suited for massively parallel workflows (maximal number of walkers, with minimal sampling time per walker) and we encourage users to exploit this.
model, walkers = learning.active_learning(
model, # used to generate hamiltonian
walkers,
steps=2000, # no more 'step' argument!
**optional_sampling_kwargs,
)
restarting a run
Learning has first-class support for restarted runs -- simply resubmit your calculation!
It will detect whether or not the corresponding output folder has already fully logged the
each of the iterations, and if so, load the final state of the model, the walkers, and the
learning instance without actually doing any calculations.